Why this unit
Ultrastructure is the operational visual language of connectomics annotation and quality control.
Technical scope
This unit focuses on EM-visible structural evidence used for compartment identity and synapse interpretation in connectomics workflows. It does not attempt full molecular cell-biology coverage; it emphasizes reproducible decisions under real proofreading constraints.
Learning goals
- Identify core neuronal ultrastructural features in EM data.
- Apply context-aware interpretation with explicit uncertainty.
- Produce annotation decisions that can be audited by a second reviewer.
Capability target
Given ambiguous EM patches, learners should make compartment and synapse calls using multi-cue evidence, attach confidence labels, and justify unresolved uncertainty without overclaiming.
Core technical anchors
- Compartments: soma, dendrite, axon, bouton, spine.
- Cues: vesicles, PSD/active zone, mitochondria, ER, microtubules.
- Multi-slice context before final annotation decisions.
Concept payload (teach explicitly)
1) Compartment cues are evidence, not labels
- Technical: compartments are inferred from combined morphology plus organellar context.
- Plain language: one clue is never enough; you need multiple clues that agree.
- Misconception to prevent: “small process = axon” (size alone is unreliable).
2) Synapse interpretation is contextual
- Technical: synapse calls require membrane apposition plus vesicle/PSD context across adjacent slices.
- Plain language: if you cannot see the neighborhood, you cannot trust a single-frame synapse guess.
- Misconception to prevent: “dark contrast means synapse” (contrast alone is not specific).
3) Uncertainty is an output, not a failure
- Technical: confidence tags preserve downstream QC integrity and guide expert review queues.
- Plain language: saying “not sure yet” with reasons is better science than forcing a wrong label.
- Misconception to prevent: “every patch must end with a hard label.”
Method deep dive: compartment-level decision protocol
- Start with local geometry (diameter changes, branching pattern, cytoplasmic density).
- Add organelle evidence (microtubule organization, mitochondria morphology, vesicle fields).
- Evaluate synaptic architecture (active zone alignment, vesicle clusters, PSD profile).
- Confirm continuity across adjacent sections before committing label.
- Assign confidence tier (
high, medium, uncertain) with rationale.
Quantitative QA checkpoints
- Inter-annotator agreement on compartment labels.
- Synapse call precision/recall on a gold-standard subset.
- Uncertain-label rate by region as an indicator of dataset difficulty.
- Turnaround time per corrected ambiguity (captures workflow scalability).
Frequent failure modes
- Single-slice overconfidence:
Resolve only after short z-stack review.
- Organelle misread due to staining variability:
Use multi-cue voting instead of one-feature decisions.
- False synapse positives in noisy contrast:
Require structural context around candidate cleft.
- Label drift across long neurite paths:
Enforce periodic consistency checks during tracing.
Visual training set
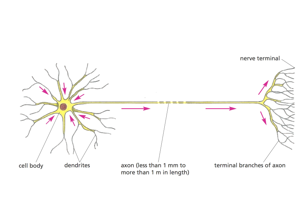
RIV-ULTRA S04: neuron-structure overview for compartment grounding.
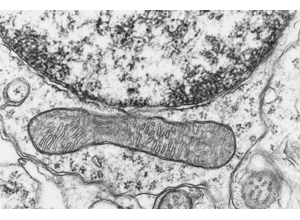
RIV-ULTRA S08: dendritic ultrastructure context.
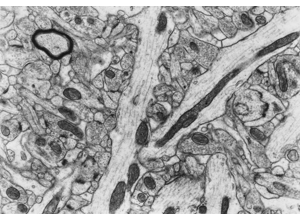
RIV-ULTRA S09: synapse-identification cue set.
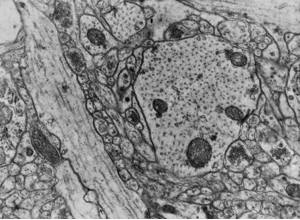
RIV-ULTRA S10: vesicle and organellar features relevant to annotation.
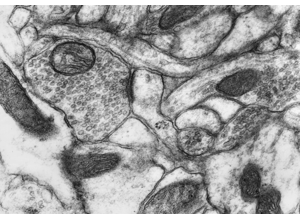
RIV-ULTRA S14: comparative ultrastructure panel.
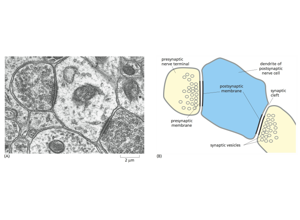
RIV-ULTRA S20: ambiguity case for context-aware interpretation.
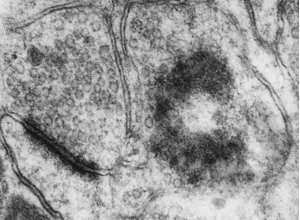
RIV-ULTRA S24: advanced structural example for review.

RIV-ULTRA S30: synthesis panel for final interpretation checks.
Attribution: Pat Rivlin training materials (MICrONS proofreading deck).
Course links
Practical workflow
- Localize candidate compartment and neighborhood context.
- Evaluate ultrastructural cues across adjacent slices.
- Assign provisional interpretation with confidence level.
- Escalate ambiguous cases for secondary review.
60-minute tutorial run-of-show (instructor-ready)
Pre-class (learner prep, 10-15 minutes async)
- Read this unit’s “Concept payload” section.
- Open the image panel below and preview at least 3 figures.
- Bring one question about a cue that seems ambiguous.
Materials needed
- Slide draft: Neuronal Ultrastructure deck draft
- Figure panel from this page (RIV-ULTRA shortlist).
- Shared annotation worksheet with columns:
- patch ID
- compartment call
- synapse call
- confidence tier
- evidence cues
Minute-by-minute plan
-
- Prompt: “What can go wrong if we force a label too early?”
- Instructor sets capability target and expected outputs.
-
| **05:00-12:00 |
Expert modeling** |
- Walk through one patch live:
- local geometry
- organelle cues
- synaptic context
- confidence assignment
- Think aloud explicitly about uncertainty.
-
| **12:00-20:00 |
Guided practice round 1** |
- Learners annotate 2 easier patches in pairs.
- Instructor circulates and checks cue quality, not just final labels.
-
| **20:00-30:00 |
Debrief + misconception check** |
- Compare labels publicly.
- Target misconceptions:
- single-cue overconfidence
- contrast-only synapse calls
- missing context across slices
-
| **30:00-42:00 |
Guided practice round 2 (ambiguous cases)** |
- Learners annotate 2 borderline patches independently.
- Require two supporting cues and one uncertainty statement per patch.
-
| **42:00-52:00 |
Consensus protocol** |
- Small groups reconcile disagreements using rubric rules.
- Escalate irreducible ambiguity with rationale.
-
| **52:00-58:00 |
Competency check** |
- Each learner submits one fully justified call:
- label
- confidence
- evidence chain
- one alternative considered
-
| **58:00-60:00 |
Exit ticket** |
- “One cue I trust more now, one cue I still mistrust.”
Instructor script cues
- “Show me your evidence chain before your label.”
- “Which cue would you drop first if contrast quality decreased?”
- “What would make you downgrade from medium to uncertain?”
- At 20 minutes: at least 80% of pairs cite two independent cues.
- At 42 minutes: group disagreement log distinguishes cue conflict vs missing context.
- At 58 minutes: learner can justify one call with explicit uncertainty language.
Post-class assignment (20-30 minutes)
- Annotate 3 new patches and submit:
- call + confidence
- cue rationale
- one unresolved ambiguity and escalation note
Studio activity: Ultrastructure consensus round
Format: 60-75 minutes, small groups, shared patch set.
Scenario: Your team is preparing a training-ready annotation subset for downstream segmentation QC. The subset contains borderline cases where compartment and synapse interpretation is uncertain.
Task sequence
- Independently label each patch: compartment, synapse status, confidence tier.
- Record two supporting cues and one uncertainty per patch.
- Compare labels within group and classify disagreements by type (cue conflict, context missing, vocabulary mismatch).
- Resolve what can be resolved with available context; escalate true ambiguities.
- Update one rubric rule to reduce future disagreement.
Expected outputs
- Consensus annotation sheet.
- Disagreement log with error-type counts.
- One rubric revision note with rationale.
Assessment rubric (unit-level)
- Minimum pass
- Uses at least two independent cues per call.
- Applies confidence tags consistently.
- Distinguishes unresolved ambiguity from error.
- Strong performance
- Cites context across slices, not only local texture.
- Produces clear disagreement taxonomy and escalation decisions.
- Improves rubric clarity based on observed disagreements.
- Common failure modes to flag
- Single-cue decisions presented as definitive.
- Synapse calls without neighborhood evidence.
- Missing or inconsistent confidence labeling.
Discussion prompts
- Which ultrastructural cues are most robust across annotators?
- Where should uncertainty remain explicit rather than forced to a hard label?
Quick activity
Using one training image, label at least three ultrastructural cues and state your confidence for each interpretation.
Content library references
- Soma ultrastructure — Nuclear envelope, Nissl substance, Golgi, lipofuscin
- Dendrite biology — Spine types, PSDs, microtubule organization, local translation
- Axon biology — AIS, myelinated segments, boutons, vesicle pools
- Synapse classification — Gray Type I/II, asymmetric vs symmetric
- Organelle annotation cues — Mitochondria, ER, MVBs as compartment indicators
- Myelin and nodes of Ranvier — Compact myelin, paranodal loops, incisures
- H01 human cortex — Human ultrastructure case study
Teaching slide deck
Evidence pack: papers and datasets
This unit is anchored to canonical papers and datasets used in connectomics practice. Use these as required preparation before activities.
Key papers
Key datasets
Competency checks
- Use at least two independent ultrastructure cues for each call.
- Tag and escalate ambiguous regions with documented rationale.
Capability development brief
Capability target: Interpret ultrastructural features reproducibly to distinguish compartments and synaptic context.
Required expertise
- Cellular neuroanatomist (organelle and compartment interpretation)
- Senior proofreader (decision consistency under ambiguity)
- Training lead (annotation rubric design)
Core concepts to teach
- Compartment cues: Features such as mitochondria density, microtubules, vesicle pools, and membrane morphology.
- Synaptic context: Interpreting cleft, vesicles, and postsynaptic density together rather than in isolation.
- Confidence tagging: Marking uncertain calls to prioritize expert review.
Studio activity
Ultrastructure Consensus Round - Build consistency in compartment and synapse labeling.
Independently annotate the same patches, then reconcile disagreements.
- Label compartments and synaptic features individually.
- Compare disagreement hotspots and identify ambiguous cues.
- Update rubric decision rules.
Expected outputs:
- Consensus annotations
- Rubric revision notes
Assessment artifacts
- Compartment annotation rubric with confidence levels.
- Inter-rater agreement report on a shared patch set.








